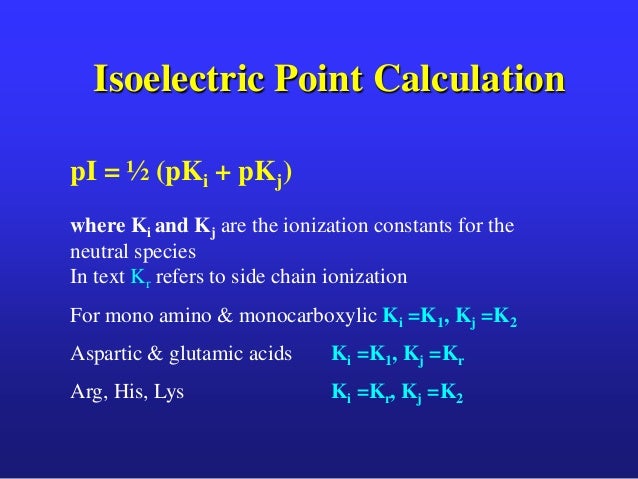

pI is most commonly examined for proteins. Overall, the net charge of the protein or peptide is strongly related to the solution (buffer) pH and can be approximated using the Henderson-Hasselbalch. Compute the theoretical pI (isoelectric point) and Mw (molecular weight) for a list of UniProt Knowledgebase (Swiss-Prot or TrEMBL) entries or for user. In general, the pI values for native proteins are affected by the three-dimensional structure of the proteins, which causes greater differences between calculated and experimental pI values than in the case of polypeptides for which pI values are determined in the presence of urea. The isoelectric point (pI) is the point at which the net charge on a molecule is zero.
The pI of the native Glut1 was lower, 8.0 +/- 0.1, at 22 degrees C. Online calculation (prediction) of theoretical isoelectric point (pI, IEP) of proteins and petides from sequence alone.

If you want to calculate pi, first measure the circumference of a circle by wrapping a piece of string around the edge of it and then measuring the length of the string.
#CALCULATE PI OF PROTEIN CODE#
The calculated pI for the human red cell glucose transporter (Glut1) with one sialic acid residue was decreased from 8.8 to 8.5 by introducing pKa value spreading and became consistent with the experimental pI value of 8.4 +/- 0.05 at 15 degrees C determined in the presence of 6 M urea. Pi is roughly 3.14, but its actually an infinite number that never slips into a repeating pattern. REM REM Revision 1.1 6 21:12:36 Don.Cross REM Batch file for calculating pi REM That is the code save it as pi.bat or anything you want. The calculated pI values showed reasonably good agreement with experimental ones for most of 16 native proteins over a wide pH range (3.4-11) when charge contributions of heme groups, sialic acid residues, etc., were taken into account. Each particular type of ionizable group was assumed to have pKa values distributed around the chosen value, thereby simulating the situation in proteins and polypeptides. A set of pKa values was chosen for amino acid residues with ionizable side chains. Amino acid composition, pKa values for amino acid side chains and for the N- and C-terminal groups, and the presence of other charged groups were taken into account. For estimating pI values the net charge of several proteins was calculated versus pH by use of the Henderson-Hasselbalch equation. At pH = 5.02, the pH = pI so the amino acid will exist as the zwitterion with both the positive and negative charges as shown above.The isoelectric points (pI) of native proteins are important in several separation techniques. Feature weights are determined from separation of low and high solubility subsets.

Using available data for Escherichia coli protein solubility in a cell-free expression system, 35 sequence-based properties are calculated. As a result, the only remaining charge will be on the carboxylate ion so the amino acid will have a \(-1\) charge.Į. ProteinSol - is a web server for predicting protein solubility.
